Advanced multi-language Interface to CVODES and IDAS (%s)
Project description
About AMICI
AMICI provides a multi-language (Python, C++, Matlab) interface for the
SUNDIALS solvers
CVODES
(for ordinary differential equations) and
IDAS
(for algebraic differential equations). AMICI allows the user to read
differential equation models specified as SBML
and automatically compiles such models as .mex simulation files
(Matlab), C++ executables or Python modules.
In contrast to the (no longer maintained) sundialsTB Matlab interface, all necessary functions are transformed into native C++ code, which allows for a significantly faster simulation.
Beyond forward integration, the compiled simulation file also allows for forward sensitivity analysis, steady state sensitivity analysis and adjoint sensitivity analysis for likelihood based output functions.
The interface was designed to provide routines for efficient gradient computation in parameter estimation of biochemical reaction models but it is also applicable to a wider range of differential equation constrained optimization problems.
Features
- SBML import (see details below)
- Generation of C++ code for model simulation and sensitivity computation
- Access to and high customizability of CVODES and IDAS solver
- Python, C++, Matlab interface
- Sensitivity analysis
- forward
- steady state
- adjoint
- first- and second-order
- Pre-equilibration and pre-simulation conditions
- Support for discrete events and logical operations
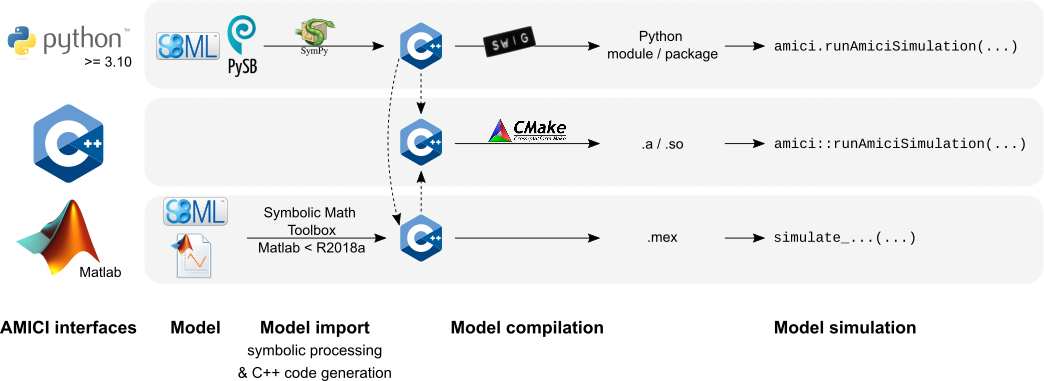
Interfaces & workflow
The AMICI workflow starts with importing a model from either
SBML (Matlab, Python) or a Matlab definition of the
model (Matlab-only). From this input, all equations for model simulation
are derived symbolically and C++ code is generated. This code is then
compiled into a C++ library, a Python module, or a Matlab .mex file and
is then used for model simulation.
Getting started
AMICI installation instructions are provided here.
To get you started with Python-AMICI the best way might be this Jupyter notebook.
For Matlab, various examples are available here.
Comprehensive documentation on installation and usage of AMICI is available online at http://icb-dcm.github.io/AMICI/.
Any contributions to AMICI are welcome, read more contributing here.
Getting help
In case of questions or problems with using AMICI, feel free to post an issue on Github. We are trying to get back to you quickly.
Publications
Citeable DOI for the latest AMICI release:
There is a list of publications using AMICI. If you used AMICI in your work, we are happy to include your project, please let us know via a Github issue.
When using AMICI in your project, please cite
- Fröhlich, F., Kaltenbacher, B., Theis, F. J., & Hasenauer, J. (2017). Scalable Parameter Estimation for Genome-Scale Biochemical Reaction Networks. Plos Computational Biology, 13(1), e1005331. doi: 10.1371/journal.pcbi.1005331 and/or
- Fröhlich, F., Theis, F. J., Rädler, J. O., & Hasenauer, J. (2017). Parameter estimation for dynamical systems with discrete events and logical operations. Bioinformatics, 33(7), 1049-1056. doi: 10.1093/bioinformatics/btw764
Status of SBML support in Python-AMICI
Python-AMICI currently passes 494 out of the 1780 (~28%) test cases from the semantic SBML Test Suite.
In additional, we currently plan to add support for the following features (see corresponding issues for details and progress):
- Events (currently Matlab-only)
- Rate rules
- Algebraic rules
- Species assignment rules
- Compartment assignment rules
- Models without species
- Logical operators
contributions are welcome.
However, the following features are unlikely to be supported:
- SBML extensions
factorial(),ceil(),floor(), due to incompatibility with symbolic sensitivity computations- initial assignments for parameters
delay()due to missing SUNDIALS solver support
Current build status




Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.