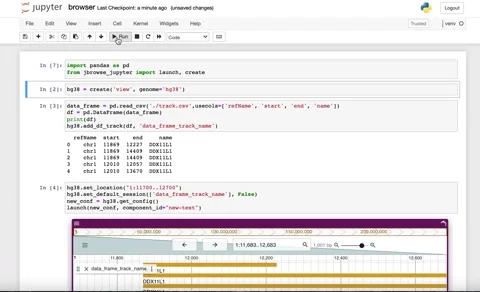
Jupyter Notebooks extension for showing JBrowse views
Project description
JBrowse Jupyter
JBrowse Jupyter is a python package that provides a python interface to JBrowse views.
The package provides a JBrowseConfig API to enable the creation of JBrowse state configuration objects. It also provides utility functions to create and embed Dash JBrowse components in jupyter notebooks and python applications.
You can open this browser.ipynb in colab here Note: to install the package in colab you can use this command. In a new cell above the import statement
!pip install git+https://github.com/GMOD/jbrowse-jupyter.git@main
Dash JBrowse
Dash JBrowse is a collection of dash components for JBrowse's embeddable components.
We utilize the Dash JBrowse package along with jupyter-dash library to embed JBrowse React Linear Genome view in any jupyter notebook.
You can find more information about our Dash JBrowse library here.
Installation
Pre-requisites
- conda - for virtual environments
- pip - python package manager
- Python - 3.6 or greater
- Jupyter - jupyter lab or jupyter notebook to run .ipynb files
PyPI
$ pip install jbrowse-jupyter
Install with conda
Clone this repository and install conda. Once you have conda installed, follow the steps found below to create a conda envirnment and install the dependecies.
$ cd jbrowse-jupyter
$ conda create -n myenv python=3.6
$ conda activate myenv
$ pip install -r requirements.txt
Quickstart
Install the JBrowse Jupyter package using pip
$ pip install jbrowse-jupyter
Launching a Linear Genome View in Jupyter Notebook
JBrowse Linear Genome view in python Dash application
import dash
import dash_html_components as html
from jbrowse_jupyter import create, create_component
app = dash.Dash(__name__)
jbrowse_conf = create("view", genome="hg38")
config = jbrowse_conf.get_config()
component = create_component(config)
app.layout = html.Div(
[component],
id='test'
)
if __name__ == "__main__":
app.run_server(port=8081, debug=True)

jbrowse_conf. The jbrowse_conf is an instance of our JBrowseConfig, and can be modified to set an assembly, add tracks, set custom color palettes and more.
Find more information about the JBrowseConfig API here
Other Examples
You can find two examples in the root of this repo,
browser.py or the browser.ipynb
browser.py uses the Dash library to create a python application with the Dash JBrowse LinearGenomeView component and configured with the help of this package. browser.ipynb is jupyter notebook using the JupyterDash library to embed a Dash JBrowse LinearGenomeView component in a cell.
To run the Python Dash application
$ python browser.py
To run the jupyter notebook
Make sure you have jupyter notebook or jupyterlab.
Within the myenv conda environment
$ pip install jupyterlab
More information on using a specific environment in a jupyter notebook
Usage
The jbrowse-jupyter package provides several utility functions to create and launch Dash JBrowse components in python applications and jupyter notebooks.
Configuring Components
create(view_type, **kwargs)- creates a JBrowseConfig configuration object given a view_type- view : creates a default empty JBrowseConfig. e.g
create('view', genome='hg19') - conf: creates a JBrowseConfig from one of the default genomes, e.g
create('conf')
- view : creates a default empty JBrowseConfig. e.g
create_component(conf, **kwargs) - creates and returns a Dash JBrowse LinearGenomeView component- component can be used as any Dash component in Dash applications
launch(conf, **kwargs) - launches a LinearGenomeView Dash component in a Jupyter cellWarning: Only use
launchin jupyter notebooks
JBrowseConfig
Quick overview of the JBrowseConfig API
The JBrowseConfig API allows us to set an assembly, add tracks, set default sessions, set custom color themes, and more.
JBrowseConfig().
-
set_assembly(assembly_data, aliases, refname_aliases)- Sets the assembly subconfiguration
- for the refname_alias subconfiguration check out the JBrowse refname aliasing docs
-
add_df_track(track_data, name, **kwargs)- requires DataFrame to have columns ['refName', 'start', 'end', 'name']
- refName and name columns must be strings, start and end must be int
- if the score column is present, it will create a Quantitative track else it will create a Feature track.
- params:
- df – pandas DataFrame with the track data.
- name (str) – (optional) name for the track
- overwrite (str) – (optional) flag wether or not to overwrite existing track.
- track_id (str) - (optional) trackId for the track
-
add_track(data, **kwargs)- adds a track configuration given a file with track data
- currently supporting cram, bam, vcf gzipped, gff/gff3 gzipped, bigbed, bigwig file types.
- assumes an index exists within the same directory of the track data if no index url path is provided.
- currently supporting Wiggle, Variant, Feature and Alignments tracks
- params:
- data (str) – track file or url (currently only supporting url)
- name (str) – (optional) name for the track
- index (str) – (optional) index file for the track
- track_type (str) – (optional) track type
- overwrite (boolean) – (optional) overwrites existing track if it exists in list of tracks (default False)
-
set_location(location)- initial location for when the browser first loads, syntax 'refName:start-end'
- e.g 'chrom1:500-1000'
-
set_default_session(tracks_ids, display_assembly=True)- sets the default session given a list of track ids
- params:
- tracks_ids - list[str] list of track ids to display
- display_assembly (boolean) – display the assembly reference sequence track. default=True
-
set_theme(primary, secondary=None, tertiary=None, quaternary=None)- sets the theme in the configuration given up to 4 hexadecimal colors
-
add_text_search_adapter(ix_path, ixx_path, meta_path, adapter_id=None)- adds a trix text search adapter
-
get_config() - returns the configuration object
DOCUMENTATION with more details and tutorials.
Resources
- JBrowse - the next generation genome browser
- JBrowse React Linear Genome View - interactive genome browser
- JBrowse React Linear Genome View Docs - storybook docs of React LGV
- Dash Applications how to get started to custumize Dash applications.
- Dash HTML components Dash html components to build the Dash aplication layout.
- DashJupyter library to enable embedding Dash components in jupyter notebooks.
- DashJbrowse suite of Dash components for JBrowse views. (JBrowse Linear Genome View)
Academic Use
This package was written with funding from the NHGRI as part of the JBrowse project. If you use it in an academic project that you publish, please cite the most recent JBrowse paper, which will be linked from jbrowse.org.
Contact us
We really love talking to our users. Please reach out with any thoughts you have on what we are building!
- Report a bug or request a feature at https://github.com/GMOD/dash_jbrowse/issues/new
- Join our developers chat at https://gitter.im/GMOD/jbrowse2
- Send an email to our mailing list at
gmod-ajax@lists.sourceforge.net
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distributions
Hashes for jbrowse_jupyter-1.1.1-py3-none-any.whl
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 | e2b0ae867c7dbb96d20dcfcff1755dff76516c266c64065c536b390cf7b3b179 |
|
| MD5 | 54b3168c163d99da3d6d76fb487182db |
|
| BLAKE2b-256 | 009829b24240ff56b7f1d47436443f7a6295f3f577e88b2f94f63f2f51a1065a |