LazyGrid: Automatic, efficient and flexible implementation of complex machine learning pipeline generation and cross-validation.
Project description
LazyGrid is a python package providing an automatic, efficient and flexible implementation of complex machine learning pipeline generation and cross-validation.
Before fitting a model or a pipeline step, LazyGrid checks inside an internal SQLite database if the model has already been fitted. If the model is found, it won’t be fitted again.
Documentation for the latest stable version is available on ReadTheDocs.
Table Of Contents
Getting Started
You can install LazyGrid along with all its dependencies from PyPI:
$ pip install -r requirements.txt lazygridor from source code:
$ git clone https://github.com/glubbdubdrib/lazygrid.git
$ cd ./lazygrid
$ pip install -r requirements.txt .LazyGrid is known to be working on Python 3.5 and above. The package is compatible with scikit-learn 0.21, tensorflow 1.14 and Keras 2.2.4.
Documentation
Documentation for the latest stable version is available on ReadTheDocs.
How to use
LazyGrid has three main features:
it can generate all possible pipelines given a set of steps (Pipeline generation) or all possible models given a grid of parameters (Grid search)
it can compare the performance of a list of models using cross-validation and statistical tests (Model comparison), and
it follows the memoization paradigm, avoiding fitting a model or a pipeline step twice.
The package is highly customizable according to the user’s needs.
Model generation
Pipeline generation
In order to generate all possible pipelines given a set of steps, you should define a list of elements, which in turn are lists of pipeline steps, i.e. preprocessors, feature selectors, classifiers, etc. Each step could be either a sklearn object or a keras model.
Once you have defined the pipeline elements, the generate_grid method will return a list of models of type sklearn.Pipeline.
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.feature_selection import SelectKBest, f_classif
from sklearn.preprocessing import RobustScaler, StandardScaler
import lazygrid as lg
preprocessors = [StandardScaler(), RobustScaler()]
feature_selectors = [SelectKBest(score_func=f_classif, k=1), SelectKBest(score_func=f_classif, k=2)]
classifiers = [RandomForestClassifier(random_state=42), SVC(random_state=42)]
elements = [preprocessors, feature_selectors, classifiers]
list_of_models = lg.grid.generate_grid(elements)Grid search
LazyGrid implements a useful functionality to emulate the grid search algorithm by generating all possible models given the model structure and its parameters.
In this case, you should define a dictionary of arguments for the model constructor and a dictionary of arguments for the fit method. The generate_grid_search method will return the list of all possible models.
The following example illustrates how to use this functionality to compare keras models with different optimizers and fit parameters.
import keras
from keras import Sequential
from keras.layers import Conv2D, MaxPooling2D, Flatten, Dense
from keras.utils import to_categorical
from sklearn.metrics import f1_score
from sklearn.datasets import load_digits
from sklearn.model_selection import StratifiedKFold
import lazygrid as lg
import numpy as np
from keras.wrappers.scikit_learn import KerasClassifier
# define keras model generator
def create_keras_model(optimizer):
kmodel = Sequential()
kmodel.add(Conv2D(32, kernel_size=(5, 5), strides=(1, 1),
activation='relu',
input_shape=x_train.shape[1:]))
kmodel.add(MaxPooling2D(pool_size=(2, 2)))
kmodel.add(Flatten())
kmodel.add(Dense(1000, activation='relu'))
kmodel.add(Dense(n_classes, activation='softmax'))
kmodel.compile(loss=keras.losses.categorical_crossentropy,
optimizer=optimizer,
metrics=['accuracy'])
return kmodel
# load data set
x, y = load_digits(return_X_y=True)
skf = StratifiedKFold(n_splits=10, shuffle=True, random_state=42)
list_of_splits = [split for split in skf.split(x, y)]
train_index, val_index = list_of_splits[0]
x_train, x_val = x[train_index], x[val_index]
y_train, y_val = y[train_index], y[val_index]
x_train = np.reshape(x_train, (x_train.shape[0], 8, 8, 1))
x_val = np.reshape(x_val, (x_val.shape[0], 8, 8, 1))
n_classes = len(np.unique(y_train))
if n_classes > 2:
y_train = to_categorical(y_train)
y_val = to_categorical(y_val)
# cast keras model into sklearn model
kmodel = KerasClassifier(create_keras_model, verbose=1, epochs=0)
# define all possible model parameters of the grid
model_params = {"optimizer": ['SGD', 'RMSprop']}
fit_params = {"epochs": [5, 10, 20], "batch_size": [10, 20]}
# generate all possible models given the parameters' grid
models, fit_parameters = lg.grid.generate_grid_search(kmodel, model_params, fit_params)You will find the conclusion of this example in the plot section.
Model comparison
Statistical hypothesis tests
Once you have generated a list of models (or pipelines), LazyGrid provides friendly APIs to compare models’ performances by using a cross-validation procedure and by analyzing the outcomes applying statistical hypothesis tests.
First, you should define a classification task (e.g. x, y = make_classification(random_state=42)), define the set of models you would like to compare (e.g. model1 = LogisticRegression(random_state=42)), and call for each model the cross_val_score method provided by sklearn.
Finally, you can collect the cross-validation scores into a single list and call the find_best_solution method provided by LazyGrid. Such method applies the following algorithm: it looks for the model having the highest mean value over its cross-validation scores (“the best model”); it compares the distribution of the scores of each model against the distribution of the scores of the best model applying a statistical hypothesis test.
You can customize the comparison by modifying the statistical hypothesis test (it should be compatible with scipy.stats) or the significance level for the test.
from sklearn.linear_model import LogisticRegression, RidgeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.datasets import make_classification
from sklearn.model_selection import cross_val_score
import lazygrid as lg
from scipy.stats import mannwhitneyu
x, y = make_classification(random_state=42)
model1 = LogisticRegression(random_state=42)
model2 = RandomForestClassifier(random_state=42)
model3 = RidgeClassifier(random_state=42)
score1 = cross_val_score(estimator=model1, X=x, y=y, cv=10)
score2 = cross_val_score(estimator=model2, X=x, y=y, cv=10)
score3 = cross_val_score(estimator=model3, X=x, y=y, cv=10)
scores = [score1, score2, score3]
best_idx, best_solutions_idx, pvalues = lg.statistics.find_best_solution(scores,
test=mannwhitneyu,
alpha=0.05)Optimized cross-validation
LazyGrid includes an optimized implementation of cross-validation (cross_validation), specifically devised when a huge number of machine learning pipelines need to be compared.
In fact, once a pipeline step has been fitted, LazyGrid saves the fitted model into a SQLite database. Therefore, should the step be required by another pipeline, LazyGrid fetches the model that has already been fitted from the database.
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.feature_selection import SelectKBest, f_classif
from sklearn.preprocessing import RobustScaler, StandardScaler
from sklearn.datasets import make_classification
import lazygrid as lg
x, y = make_classification(random_state=42)
preprocessors = [StandardScaler(), RobustScaler()]
feature_selectors = [SelectKBest(score_func=f_classif, k=1),
SelectKBest(score_func=f_classif, k=2)]
classifiers = [RandomForestClassifier(random_state=42), SVC(random_state=42)]
elements = [preprocessors, feature_selectors, classifiers]
models = lg.grid.generate_grid(elements)
for model in models:
model = lg.wrapper.SklearnWrapper(model, dataset_id=1, db_name="./database/sklearn-db.sqlite",
dataset_name="make-classification")
score, fitted_models, \
y_pred_list, y_true_list = lg.model_selection.cross_validation(model=model, x=x, y=y)Automatic reports
The compare_models method provides a friendly approach to compare a list of models: it calls the cross_validation method for each model, automatically performing the optimized cross-validation using the memoization paradigm; it calls the find_best_solution method, applying a statistical test on the cross-validation results; it returns a Pandas.DataFrame containing a summary of the results.
from sklearn.linear_model import LogisticRegression, RidgeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.datasets import make_classification
import pandas as pd
import lazygrid as lg
x, y = make_classification(random_state=42)
lg_model_1 = lg.wrapper.SklearnWrapper(LogisticRegression())
lg_model_2 = lg.wrapper.SklearnWrapper(RandomForestClassifier())
lg_model_3 = lg.wrapper.SklearnWrapper(RidgeClassifier())
models = [lg_model_1, lg_model_2, lg_model_3]
results = lg.model_selection.compare_models(models=models, x_train=x, y_train=y)Utilities
Customize your cross-validation score
By default, during the cross-validation procedure, LazyGrid exploits as score
function the built-in score method of the current model, calling
model.score(x, y).
However, two levels of customization are provided. The first one allows you to use custom sklear-like score functions (e.g. accuracy_score or f1_score). You just need to call the cross-validation procedure specifying the desired score function:
import lazygrid as lg
from sklearn.metrics import f1_score
...
lg.model_selection.cross_validation(model, x, y, score_fun=f1_score)Alternatively, if you really need something different, you could write your own score function. LazyGrid assigns to the generic_score method all available local variables at each cross-validation step, giving you maximum power and flexibility:
...
score[split_index] = generic_score(**locals())
...As an example, you could use a score function to measure the class-imbalance ratio of the validation set:
import numpy as np
def compute_class_imbalance_ratio(y_val, *args, **kwargs):
"""
Compute class-imbalance ratio of the validation set.
"""
values, counts = np.unique(y_val, return_counts=True)
pmax = np.max(counts) # majority class
pmin = np.min(counts) # minority class
imbalance_ratio = pmax / pmin
return imbalance_ratioand use it when calling the cross-validation procedure:
import lazygrid as lg
...
lg.model_selection.cross_validation(model, x, y, generic_score=compute_class_imbalance_ratio)Customize your Wrapper
LazyGrid provides several classes to wrap machine learning models to make them able to interface properly with a SQLite database where fitted models will be stored. In order to use LazyGrid methods you should wrap your models first. Model wrappers include classes as: SklearnWrapper, PipelineWrapper (for sklearn pipelines), and KerasWrapper.
Moreover you can extend the abstract class Wrapper and customize the wrapper behavior according to your needs. You just need to implement the set_random_seed and the parse_parameters abstract methods. The easiest (but deprecated) way could be skipping them as follows:
from lazygrid.wrapper import Wrapper
class CustomWrapper(Wrapper):
def __init__(self, **kwargs):
Wrapper.__init__(self, **kwargs)
def set_random_seed(self, seed, split_index, random_model, **kwargs):
pass
def parse_parameters(self, **kwargs) -> str:
passPlot your results
Should you need a visual output of the results, LazyGrid includes the generate_confusion_matrix method to save a cunfusion matrix figure and to return a pycm ConfusionMatrix object.
The following lines conclude the keras example:
...
# define scoring function for one-hot-encoded lables
def score_fun(y, y_pred):
y = np.argmax(y, axis=1)
y_pred = np.argmax(y_pred, axis=1)
return f1_score(y, y_pred, average="weighted")
db_name = "./database/database.sqlite"
dataset_id = 2
dataset_name = "digits"
# cross validation
for model, fp in zip(models, fit_parameters):
model = lg.wrapper.KerasWrapper(model, fit_params=fp, db_name=db_name,
dataset_id=dataset_id, dataset_name=dataset_name)
score, fitted_models, \
y_pred_list, y_true_list = lg.model_selection.cross_validation(model=model, x=x_train, y=y_train,
x_val=x_val, y_val=y_val,
random_data=False, n_splits=3,
scoring=score_fun)
conf_mat = lg.plotter.generate_confusion_matrix(fitted_models[-1].model_id, fitted_models[-1].model_name,
y_pred_list, y_true_list, encoding="one-hot")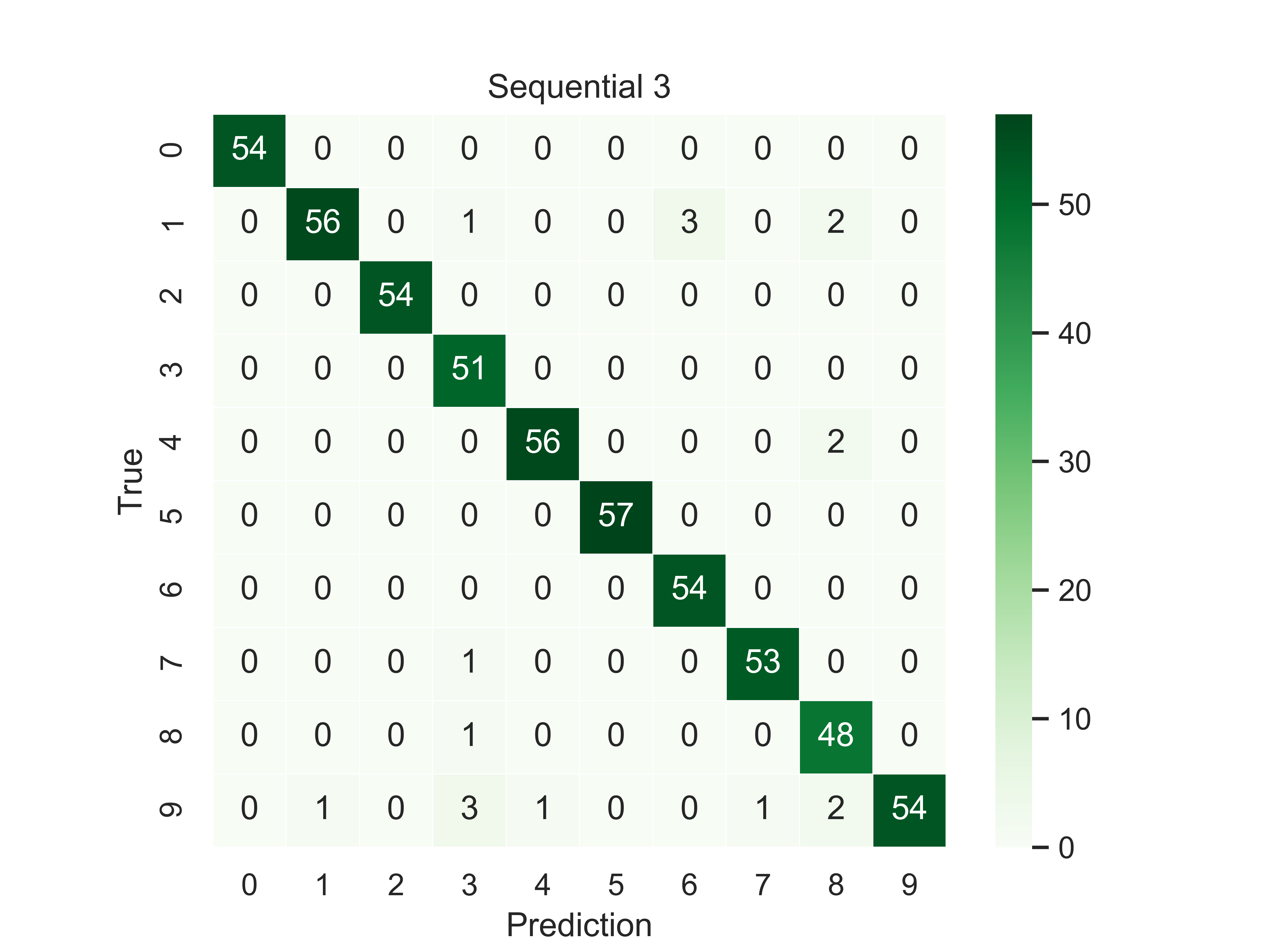
If you are looking for a visual representation of your cross-validation scores, you may use the ``
from sklearn.linear_model import LogisticRegression, RidgeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.datasets import make_classification
import lazygrid as lg
x, y = make_classification(random_state=42)
lg_model_1 = lg.wrapper.SklearnWrapper(LogisticRegression())
lg_model_2 = lg.wrapper.SklearnWrapper(RandomForestClassifier())
lg_model_3 = lg.wrapper.SklearnWrapper(RidgeClassifier())
models = [lg_model_1, lg_model_2, lg_model_3]
score_list = []
labels = []
for model in models:
scores, _, _, _ = lg.model_selection.cross_validation(model, x, y)
score_list.append(scores["val_cv"])
labels.append(model.model_name)
file_name = "val_scores"
title = "Model comparison"
lg.plotter.plot_boxplots(score_list, labels, file_name, title)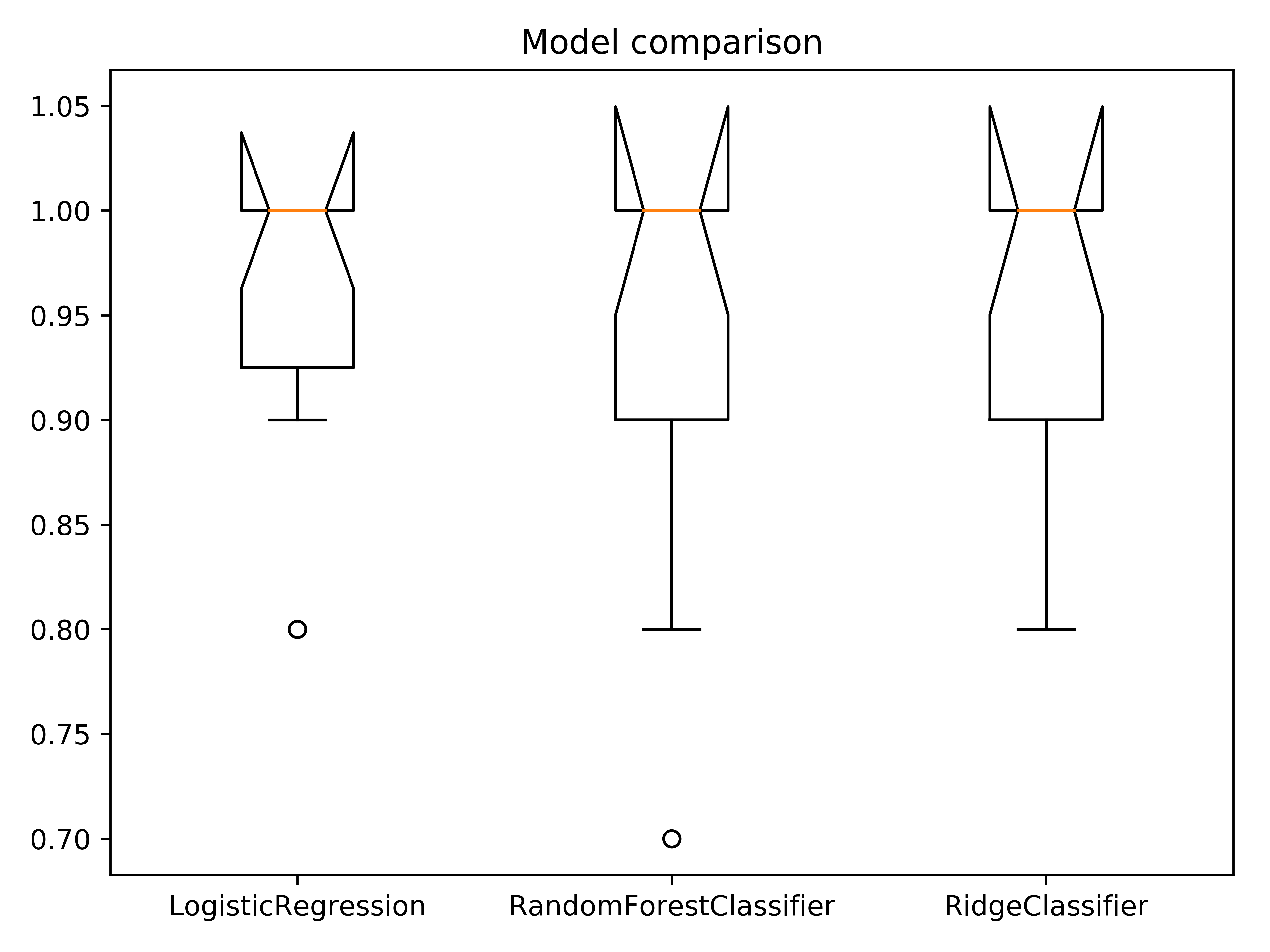
Data set APIs
LazyGrid includes a set of easy-to-use APIs to fetch OpenML data sets (NB: OpenML has a database of more than 20000 data sets).
The fetch_datasets method allows you to smartly handle such data sets: it looks for OpenML data sets compliant with the requirements specified; for such data sets, it fetches the characteristics of their latest version; it saves in a local cache file the properties of such data sets, so that experiments can be easily reproduced using the same data sets and versions. You will find the list of downloaded data sets inside ./data/<datetime>-datalist.csv.
The load_openml_dataset method can then be used to download the required data set version.
import lazygrid as lg
datasets = lg.datasets.fetch_datasets(task="classification", min_classes=2,
max_samples=1000, max_features=10)
# get the latest (or cached) version of the iris data set
data_id = datasets.loc["iris"].did
x, y, n_classes = lg.datasets.load_openml_dataset(data_id)Running tests
You can run all unittests from command line by using python:
$ python -m unittest discoveror coverage:
$ coverage run -m unittest discoverContributing
Please read Contributing.md for details on our code of conduct, and the process for submitting pull requests to us.
Licence
Copyright 2019 Pietro Barbiero and Giovanni Squillero.
Licensed under the Apache License, Version 2.0 (the “License”); you may not use this file except in compliance with the License. You may obtain a copy of the License at: http://www.apache.org/licenses/LICENSE-2.0.
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an “AS IS” BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and limitations under the License.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.


























