NGS, ribosomal, rRNA, snakemake, sequana
Project description



This is is the ribofinder pipeline from the Sequana project
- Overview:
Simple parallele workflow to detect and report ribosomal content
- Input:
FastQ files
- Output:
HTML reports
- Status:
production
- Citation:
Cokelaer et al, (2017), ‘Sequana’: a Set of Snakemake NGS pipelines, Journal of Open Source Software, 2(16), 352, JOSS DOI doi:10.21105/joss.00352
Installation
You must install Sequana first (use –upgrade to get the latest version installed):
pip install sequana --upgrade
Then, just install this package:
pip install sequana_ribofinder --upgrade
Usage
This command will scan all files ending in .fastq.gz found in the local directory, create a directory called fastqc/ where a snakemake pipeline is launched automatically. Depending on the number of files and their sizes, the process may be long:
sequana_pipelines_ribofinder --help sequana_pipelines_ribofinder --input-directory DATAPATH
This creates a directory with the pipeline and configuration file. You will then need to execute the pipeline:
cd ribofinder sh ribofinder.sh # for a local run
This launch a snakemake pipeline. If you are familiar with snakemake, you can retrieve the pipeline itself and its configuration files and then execute the pipeline yourself with specific parameters:
snakemake -s ribofinder.rules -c config.yaml --cores 4 --stats stats.txt
Or use sequanix interface.
Requirements
This pipelines requires the following executable(s):
bowtie1
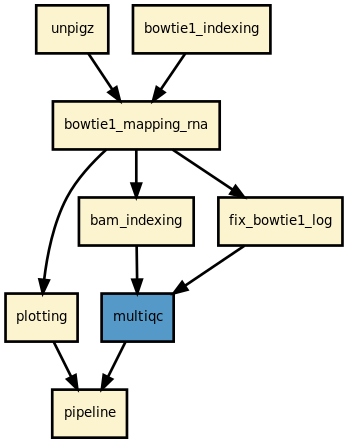
Details
This pipeline runs ribofinder in parallel on the input fastq files. A brief sequana summary report is also produced.
Rules and configuration details
Here is the latest documented configuration file to be used with the pipeline. Each rule used in the pipeline may have a section in the configuration file.
Changelog
Version |
Description |
|---|---|
0.10.1 |
|
0.10.0 |
|
0.9.3 |
|
0.9.2 |
First release. |
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Hashes for sequana_ribofinder-0.10.1.tar.gz
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 | 7fafb3d32cdf39acacb0dabd09560cd2ed4d37a52dcfd473c187f94cf95574a1 |
|
| MD5 | 429b127a60733613c72fdefd3201f9d5 |
|
| BLAKE2b-256 | 8fcff78a0ec2c09c8166d27a8614870de6a1a371fd1e7102783857195ed01195 |











