Research experimental pipeline automation for multi-agent simulations and real robots
Project description
SIERRA (reSearch pIpEline for Reproducability, Reusability, and Automation)
| Usage | Release | Development |
|---|---|---|
 |
 |
 |
 |
 |
 |
  |
 |
 |
 |
 |
 |
Installing SIERRA
pip3 install sierra-research
Why SIERRA?
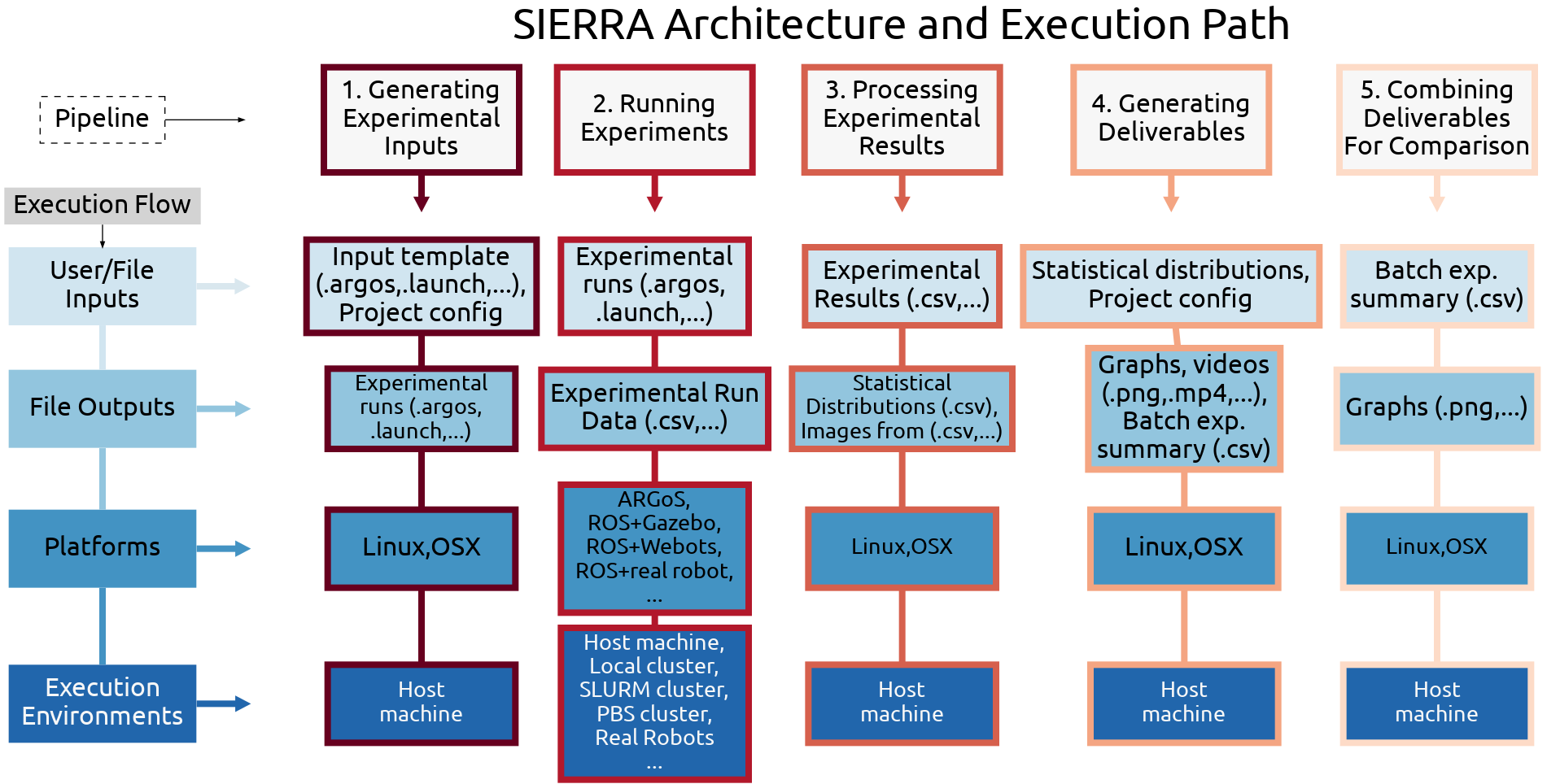
SIERRA is a command line tool for automating the pipeline described above, providing faculties for seamless experiment generation, execution, and results processing. Essentially, SIERRA handles the “backend” parts of research, such as: handling random seeds, algorithm stochasticity, configuration for a given execution environment or platform, generating statistics from experimental results, and generating graphs and videos from experimental results. This allows researchers to focus on the “research” aspects: developing theories and algorithms and testing them via experimental evaluation. SIERRA also eliminates menial reconfiguration of experimental inputs across platforms and execution environments by decoupling the two concepts: any supported execution environment or platform can be selected with a single command line switch. It further automates the process of experimental data processing and deliverable generation, changing the paradigm of those pipeline stages from procedural (“Do these steps in order to process my data and generate graphs”) to declarative (“Here are the deliverables I want to generate and the data I want to appear on them”), eliminating the need for throw-away scripts. Overall, SIERRA’s automation increases reusability and reproducability across projects, which lowers the barriers to collaboration between researchers, while also accelerating research and development cycles.
Consider the two use cases below: within each, SIERRA provides faculties for managing heterogeneity and automating common tasks to reduce the burden on researchers, either directly or through a rich plugin interface. If aspects of either use case sound familiar, then there is a strong chance SIERRA could help you with your research! SIERRA is well documented--see the docs here for more info on the automation it provides and to get started using it!
Use Case #1: Alice The Robotics Researcher
Alice is a researcher at a large university that has developed a distributed new task allocation algorithm $\alpha$ for use in a foraging task where robots must coordinate to find objects of interest in an unknown environment and bring them to a central location. Alice wants to implement her algorithm so she can investigate:
-
How well it scales with the number of robots, specifically if it remains efficient with up to 1000 robots in several different scenarios.
-
How robust it is with respect to sensor and actuator noise.
-
How it compares to other similar state of the art algorithms on a foraging task: $\beta,\gamma$.
Alice is faced with the following heterogeneity matrix which she has to deal with in addition to theoretical and algorithmic issues in her algorithms of interest in order to answer her research queries:
| Algorithm | Contains stochasticity? | Outputs data in? |
|---|---|---|
| $\alpha$ | Yes | .csv, .rosbag |
| $\beta$ | Yes | .csv, .rosbag |
| $\gamma$ | No | .rosbag |
Alice is familiar with ROS, and wants to use it with large scale simulated and small scale real-robot experiments with TurtleBots. However, for real robots she is unsure what data she will ultimately need, and wants to capture all ROS messages, to avoid having to redo experiments later. She has access to a large SLURM-managed cluster, and prefers to develop code on her laptop.
Use Case#2: Alice The Contagion Modeler
Alice has teamed with Bob, a biologist, to model the spread of contagion among agents in a population, and how that affects their individual and collective abilities to do tasks. She believes her α algorithm can be reused in this context. However, Bob is not convinced and has selected several multi-agent models from recent papers: $\delta$,$\epsilon$, and wants Alice to compare $\alpha$ to them. $\delta$ was originally developed in NetLogo, for modeling disease transmission in animals. $\epsilon$ was originally developed for ARGoS to model the effects of radiation on robots.
Alice is faced with the following heterogeneity matrix which she must deal with with in addition to theoretical and algorithmic issues in her algorithms of interest in order to answer her research queries:
| Algorithm | Can Run On? | Input Requirements? |
|---|---|---|
| $\alpha$ | ROS/Gazebo | XML |
| $\delta$ | NetLogo | NetLogo |
| $\epsilon$ | ARGoS | XML |
Bob is interested in how the rate of contagion spread varies with agent velocity and population size. Bob needs to prepare succinct, comprehensive visual representations of the results of this research queries for a a presentation, including visual comparisons of the multi-agent model as it runs for each algorithm. He will give Alice a range of parameter values to test for each algorithm based on his ecological knowledge, and rely on Alice to perform the experiments. For this project, Alice does not have access to HPC resources, but does have a handful of servers in her lab which she can use.
SIERRA Support Matrix
SIERRA supports multiple platforms which researchers can write code to target (docs). SIERRA supports multiple execution environments for execution of experiments, such as High Performance Computing (HPC) environments (docs) and real robots (docs). If your desired platform or execution environment is not listed, see the docs for how to add it via a plugin.
| Execution Environment | Description |
|---|---|
| SLURM | A cluster managed by the SLURM scheduler |
| Torque/MOAB | A cluster managed by the Torque/MOAB scheduler |
| ADHOC | Miscellaneous collection of networked compute nodes (not managed by a scheduler) |
| Local | The SIERRA host machine (e.g., a researcher's laptop) |
| Turtlebot3 | Real turtlebot3 robots |
| Platform | Description |
|---|---|
| ARGoS | Simulator for fast simulation of large swarms. Requires ARGoS >= 3.0.0-beta59. |
| ROS1+Gazebo | Using ROS1 with the Gazebo simulator. Requires Gazebo >= 11.9.0, ROS1 Noetic or later. |
| ROS1+Robot | Using ROS1 with a real robot platform of your choice. ROS1 Noetic or later is required. |
SIERRA also supports multiple output formats for experimental outputs. If the format for your experimental outputs is not listed, see the docs for how to add it via a plugin. SIERRA currently only supports XML experimental inputs.
| Experimental Output Format | Scope |
|---|---|
| CSV file | Raw experimental outputs, tranforming into heatmap images |
| PNG file | Stitching images together into videos |
SIERRA supports (mostly) mix-and-match between platforms, execution environments, experiment input/output formats as shown in its support matrix below. This is one of the most powerful features of SIERRA!
| Execution Environment | Platform | Experimental Input Format | Experimental Output Format |
|---|---|---|---|
| SLURM | ARGoS, ROS1+Gazebo | XML | CSV, PNG |
| Torque/MOAB | ARGoS, ROS1+Gazebo | XML | CSV, PNG |
| ADHOC | ARGoS, ROS1+Gazebo | XML | CSV, PNG |
| Local | ARGoS, ROS1+Gazebo | XML | CSV, PNG |
| ROS1+Turtlebot3 | ROS1+Gazebo, ROS1+robot | XML | CSV, PNG |
Requirements To Use SIERRA
The basic requirements are:
-
Recent OSX or Linux (Windows is not supported).
-
python >= 3.9.
For more details, such requirements for researcher code, see the docs.
Citing
If you use SIERRA and have found it helpful, please cite the following paper:
@inproceedings{Harwell2022a-SIERRA,
author = {Harwell, John and Lowmanstone, London and Gini, Maria},
title = {SIERRA: A Modular Framework for Research Automation},
year = {2022},
isbn = {9781450392136},
publisher = {International Foundation for Autonomous Agents and Multiagent Systems},
booktitle = {Proceedings of the 21st International Conference on Autonomous Agents and Multiagent Systems},
pages = {1905–1907}
}
Contributing
See here to get started.
License
This project is licensed under GPL 3.0. See LICENSE.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Hashes for sierra_research-1.2.7-py3-none-any.whl
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 | b00c01feb8debd917fa443f5a54a1146057edc54b390f287612e46066992c436 |
|
| MD5 | 9fbbbbf6802dc02f5b3ebe7030e54b43 |
|
| BLAKE2b-256 | 678537dc3b0e4fecfe57a8f31c5d694b69c601ea21e48c1b5a44aa39926e9d8e |