Python implementation of the kinetic model of neuromuscular transmission dynamics.
Project description
Dynamics of Neuromuscular Transmission Reproduced by Calcium-Dependent and Reversible Serial Transitions in the Vesicle Fusion Complex
KiNeuron: Python implementation of the kinetic model of neuromuscular transmission dynamics.
KiNeuron is an open-source implementation of our mechanistic kinetic model of neuromuscular transmission based on sequential maturation transitions in the molecular fusion complex.
KiNeuron is:
- Simple -- It is possible to simulate an alternative kinetic model with a few lines of code. This way, you can focus on the key parts of the problem that really matter.
- Flexible -- It is possible to customise the kinetic model by adjusting the number of transition states, the kinetic transitions between states, as well as the rate constants. Further, KiNeuron allows the addition of a stimulation protocol.
- Source Code: https://github.com/alexini-mv/kinetic-neurotransmission
- Bug reports: https://github.com/alexini-mv/kinetic-neurotransmission/issues
- Citing in your work: https://doi.org/10.3389/fnsyn.2021.785361
Requirements
KiNeuron requires:
- python >= 3.6
- graphviz >= 0.19.1
- matplotlib >= 3.3.4
- numpy >= 1.19.5
- pandas >= 1.1.5, <1.3.0
To use the graph display functions of the model, it is necessary to install the Graphviz library as described in following documentation.
Installation
There are two ways to install KiNeuron:
-
Via PyPI repository (recommended):
- In your local workspace, create a new Python Virtual Environment with venv or conda.
- Install KiNeuron as follow:
$ python -m pip install -U kineuron
- All dependencies are downloaded and installed. To verify that it has been installed correctly, you can check the following output:
$ python -c "import kineuron; print(kineuron.__version__)" '0.1.0'
-
Via GitHub:
- Clone this project repository to your local workspace.
- Create a new Python Virtual Environment with venv or conda.
- Install the required libraries from the file
requeriments.txtinto your virtual environment as follow:
$ python -m pip install -r requeriments.txt
You are ready to use KiNeuron.
Usage Example
Creating a Model
Create a file main.py and import the following classes:
from kineuron import (KineticModel, RateConstant, Solver, Stimulation,
Transition, TransitionState)
Then, instantiate the model objects as follows:
model = KineticModel(name='my-model', vesicles=100)
docked = TransitionState(name='Docked')
fusion = TransitionState(name='Fusion')
alpha = RateConstant(name="α", value=0.3, calcium_dependent=True)
beta = RateConstant(name="β", value=15)
tr1 = Transition(name='Transition 1',
rate_constant=alpha,
origin="Docked",
destination="Fusion")
tr2 = Transition(name='Transition 2',
rate_constant=beta,
origin="Fusion",
destination="Docked")
Add all objects to the model as follow:
model.add_transition_states([docked, fusion])
model.add_rate_constants([alpha, beta])
model.add_transitions([tr1, tr2])
Finally, initialize the model:
model.init()
Likewise, a stimulation protocol should be defined (if the experiment expects it) as follows:
protocol = Stimulation(
conditional_stimuli=5,
period=0.03,
time_start_stimulation=0.1,
tau_stimulus=0.0013,
time_wait_test=0.2,
intensity_stimulus=100.0,
type_stimulus='exponential_decay',
name="Custom Stimulation Protocol")
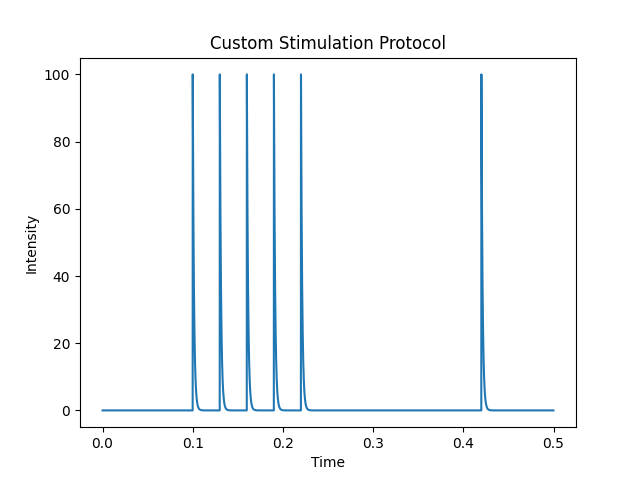
The following lines show the time profile of the stimulation protocol:
import numpy as np
t = np.arange(0, 0.5, 0.0001)
protocol.plot(t)
Model Information (Optional)
The general model information can be obtained as follows:
model.get_info()
and run the file main.py:
$ python main.py
============== MODEL INFORMATION ===============
MODEL NAME: my-model
TOTAL VESICLES: 100
TRANSITION STATES VESICLES
Docked: 100
Fusion: 0
--------------------------------------------------
NAME TRANSITION: Transition 1
RATE CONSTANT NAME: α
RATE CONSTANT VALUE: 0.3 s⁻¹
CALCIUM-DEPENDENT: True
ORIGIN: Docked
DESTINATION: Fusion
--------------------------------------------------
NAME TRANSITION: Transition 2
RATE CONSTANT NAME: β
RATE CONSTANT VALUE: 15 s⁻¹
CALCIUM-DEPENDENT: False
ORIGIN: Fusion
DESTINATION: Docked
==================================================
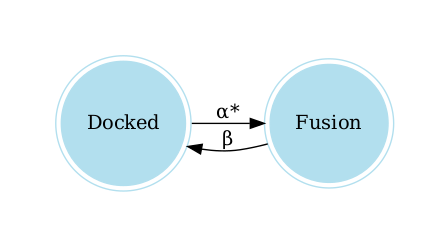
The following lines allow you to visualize the graph of the model:
graph = model.get_graph()
graph.view()
Run It
The Solver object that simulates the time evolution of the model must be instantiated. Here, we use an implementation of the Gillespie Stochastic Algorithm (1977).
experiment = Solver(model=model, stimulation=protocol)
Before initiating the simulation, make sure to obtain the resting state of the model, from which all repetitions of the experiment are starting. This is achieved as follows:
experiment.resting_state()
With the following lines the experiment is ran. The results can be obtained and saved in a .csv file for further analysis.
experiment.run(repeat=1)
results = experiment.get_results(mean=True)
results.to_csv("results.csv", index=True)
Finally, the main.py file should be executed to perform the complete simulation:
$ python main.py
Release History
- 0.1.0
- Stable version released.
- 0.0.1
- Work in progress.
Contributing
-
If you are interested in contributing to the project, please follow these guidelines:
- Fork it (https://github.com/alexini-mv/kinetic-neurotransmission).
- Create your feature branch:
$ git checkout -b feature/fooBar
- Commit your changes:
$ git commit -am 'Add some fooBar'
- Push to the branch:
$ git push origin feature/fooBar
- Create a new Pull Request.
-
If you want to report a bug, please create a new issue here describing the error as clearly as possible.
Correspondence
Author: Alejandro Martínez-Valencia
Email: al.martinez.valencia@gmail.com
Citation
If you use our code for your research or scientific publication, we kindly ask you to refer to our work as follows:
- Martínez-Valencia A., Ramírez-Santiago G. and De-Miguel F.F. (2022) Dynamics of Neuromuscular Transmission Reproduced by Calcium-Dependent and Reversible Serial Transitions in the Vesicle Fusion Complex. Front. Synaptic Neurosci. 13:785361. DOI: 10.3389/fnsyn.2021.785361
License
Distributed under the GNU General Public License. See LICENSE for more information.
Interesting Links
- Perkel, Jeffrey M. (2022) How to fix your scientific coding errors. Nature 602, 172-173. DOI: 10.1038/d41586-022-00217-0.
- Pylint, a tool that checks for errors in Python code. Tutorials and documentation.
- Autopep8, a package automatically formats Python code to conform to the PEP 8 style guide. Documentation.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.















